Biological Nutrient Removal DNA Sequencing
About the author:
John Tillotson is managing partner for WaterTrust. Tillotson can be reached at [email protected].
With most wastewater biological nutrient removal (BNR) processes, microbe identities and abundances are not directly measured or tracked because BNR microbes are very small, and not identifiable with standard industry tests. With modern molecular test methods, such as next generation DNA sequencing, it is now possible to digitize the ecology of BNR processes at an affordable price, identifying, quantifying, tracking and bio-mapping nearly all microbes present.
BNR DNA Sequencing Case Study
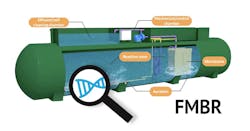
The facultative membrane bio-reactor (FMBR) is a new type of membrane bio-reactor (MBR) wastewater treatment technology that was recently documented to reduce energy costs by 77% as compared to a sequence batch reactor (SBR) that it replaced in a pilot demonstration project. It also saved 65% in biosolids disposal and had a 75% smaller footprint.
FMBR technology was invented by Jiangxi JDL Environmental Protection Co., Ltd. (JDL) of Nanchang, China in 2008. JDL claims 47 invention patents across the USA, UK, France, Japan, China, and other countries, and more than 3,000 systems installed and commissioned across 19 countries.
The FMBR is a biological process optimized to simultaneously remove carbon, nitrogen, and phosphorus from wastewater in one step, at low energy. Its design is based on proprietary environmental control strategies which select for specific types of BNR microbes without needing to add any inputs other than a continuous supply of wastewater and a low amount of electric energy to power the system. The outputs are clean water effluent that meet NPDES permit requirements or are reused on site for water conservation and sustainability benefits. The design characteristics make it an efficient decentralized wastewater treatment system (DEWATS).
FMBR – How is it different?
Some modern MBR systems complete nitrification and denitrification simultaneously, in a low dissolved oxygen (DO) condition, saving energy and footprint. Normally, phosphorus is removed in a different process. With the FMBR, phosphorus is removed biologically in the same reactor and ecological environment where simultaneous nitrification and denitrification occurs, further reducing footprint and cost.
FMBR Pilot DNA Sequencing to Identify What Bacteria Is There and How Many?
In Phase I of this study, Microbe Detectives (MD) analyzed 13 samples of the FMBR pilot, between May 2020 and May 2021. Standard MD 16S DNA sequencing methods were applied, identifying nearly all bacteria and archaea at the genus level or above and their relative percentage abundance.
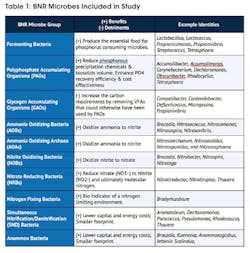
The data analysis was specialized for BNR applications with the intent to create key observations. Key BNR microbes of interest that have been shown to perform beneficial biological nutrient removal functions were identified and measured for their percent relevant abundance, summarized in Table 1.
How Do the BNR Biocharacteristics Compare?
MD 16S DNA sequencing data of the FMBR pilot samples was aggregated with MD 16S sequencing database of 675 samples from 18 municipal wastewater BNR processes that are dispersed across the New England, Midwest, Southwest, Rocky Mountains and West Coast geographies in the U.S., with daily average flows ranging from less than 1 million gallons per day (mgd) to greater than 100 mgd, summarized in Table 2. All data was anonymized. Statistical summaries of the presence and abundance of BNR microbes observed in each BNR process (“benchmark”) were compared.
Key Observations
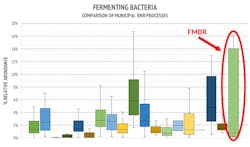
Carbon (C) Removal. Fermenting bacteria remove C by decomposing organic waste. They produce volatile fatty acids (VFAs), which are food for phosphate accumulating organisms (PAOs). PAOs remove phosphorus from wastewater biologically (BioP). The percent abundance of fermenting bacteria observed in FMBR pilot samples averaged 4.2% vs 2.4% in BNR benchmarks. Tetrasphaera represented approximately 95% of fermenters. As a denitrifying PAO (DPAO), Tetrasphaera can utilize internally stored carbon both for phosphorus uptake and denitrification at the anoxic stage, resulting in increased phosphorus and nitrogen removal. In addition, DPAOs have stronger endogenous respiration, which reduces sludge production [1].
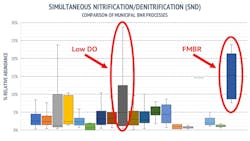
Nitrogen (N) Removal. The FMBR pilot system was mainly relying on simultaneous nitrification and denitrification (SND) bacteria to remove N. This is a unique process, which is usually operated at low dissolved oxygen concentrations (less than 0.5 mg/L), which results in aerobic conditions in the outside of the floc but anoxic conditions inside. This allows these bacteria to both oxidize ammonia aerobically and perform nitrate reduction in anoxic conditions. SND bacteria require 20 to 30% less oxygen and 40% less carbon than traditional N bacteria and have stronger endogenous respiration, which reduces sludge production.
An average 17.6% abundance of N bacteria was observed in FMBR samples vs 6.3% in BNR benchmarks, 94% of which were SND bacteria. Dechloromonas (average 8.3% in FMBR vs 1.0% in BNR benchmarks), Pseudomonas (average 8.1% in FMBR vs 3.1% in BNR benchmarks), and Tetrasphaera (average 4.0% in FMBR vs 2.4% in BNR benchmarks), were detected in all 13 FMBR samples, often at a relatively high percent abundance.
Phosphorus (P) Removal. PAOs have a unique metabolic ability to take up and store VFAs, such as acetic acid during anaerobic conditions. PAOs consume polyphosphate for energy during anaerobic conditions and take up organics and store them as polyhydroxy acetate (PHA). They then consume PHA for energy during aerobic conditions and take up phosphate to store energy as a polyphosphate granule.
PAOs were observed at an average 12.4% in FMBR Pilot samples vs 4.1% in BNR benchmarks. Tetrasphaera has been shown to be a very robust PAO and was identified in all 13 samples. In the first three samples it was observed in the the range of 14% to 16%. Overall PAO percent abundance in FMBR samples was the highest among all benchmarks.
Traditional BNR Microbes are Absent. Key microbes common in traditional BNR processes were largely absent. This included Ammonia oxidizing bacteria (AOBs) (average 0.1% in FMBR samples), nitrite oxidizing bacteria (NOBs) (average 1.0%), and nitrate reducing bacteria (NRBs) (average 0.1%). In a low dissolved oxygen SND condition, these microbes are expected to be absent because the environmental conditions are not suitable.
DNA Data & Operational Data Confirm Results. Daily influent and effluent test data of C, total nitrogen (TN), biological oxygen demand (BOD), and total suspended solids (TSS) removal by the FMBR Pilot demonstrated strong performance. MD 16S DNA sequencing confirmed the FMBR pilot system was mainly relying on SND bacteria to remove nitrogen. SND require 20 to 30% less oxygen and 40% less C than most other N bacteria. This translated into a 77% energy savings. The high abundance of SND and DPAO bacteria, which have stronger endogenous respiration, reduced sludge production by 50%. Combined with other factors, annual biosolids volume requiring offsite disposal was reduced by 65%. In the end, the DNA and operational data confirmed the results – simultaneous removal of C, N, and P, in a single tank with a surprisingly small amount of energy, footprint and biosolids waste.
References
1. Kristiansen, R., Nguyen, H., Saunders, A. et al. “A metabolic model for members of the genus Tetrasphaera involved in enhanced biological phosphorus removal.” ISME J 7, 543–554 (2013).
2. Xu, D., Liu, S., Chen, Q. et al. “Microbial community compositions in different functional zones of Carrousel oxidation ditch system for domestic wastewater treatment.” AMB Expr 7, 40 (2017). https://doi.org/10.1186/s13568-017-0336-y
3. Lifang Luo, Junqin Yao*, Weiguo Liu et al. “Comparison of bacterial communities and antibiotic resistance genes in oxidation ditches and membrane bioreactors.” Nature Portfolio, Scientific Reports (2021)
4. Jin, R., Liu, T., Liu, G. et al. “Simultaneous Heterotrophic Nitrification and Aerobic Denitrification by the Marine Origin Bacterium Pseudomonas sp. ADN-42.” Appl Biochem Biotechnol 175, 2000–2011 (2015).
5. Bergey’s Manual of Systematic Bacteriology, Edition Eight, by R.E. Buchanan and N.E. Gibbens, page 517.
6. Strous M.; et al. “Deciphering the evolution and metabolism of an anammox bacterium from a community genome.” Nature. 440 (7085): 790–794. (2006).
7. Lv, Xiao-Mei et al. “A comparative study of the bacterial community in denitrifying and traditional enhanced biological phosphorus removal processes.” Microbes and environments vol. 29(3): 261-268. (2014).